Research Progress – HIGHLIGHTS
Activity 1. Climate adaptation
?
Activity 1.1 Candidate genes in Douglas-fir
To identify the candidates genes relating to drought and frost tolerance we will establish 1) a large outdoor common garden trail to test frost hardiness, and 2) a large indoor container experiment to examine response to drought stress. We will grow trees from 60 wild collected (natural) populations and 11 orchard seedlots (selectively breed trees). Of the 60 wild seedlots, 20 will be used for the case-control analysis for genotype-phenotype association and genotype-enviromnent tests. The orchard seedlots we will be using are also part of a separate study on assisted migration. These orchard seedlots are presently growing on 45 different sites (i.e. climates) across western Canada and the US where they are being phenotyped for survial and growth.
The seedlings (approximately 10,000 individuals) will spend there first year under similar conditions in the Horticulure greenhouse at UBC. During their first growing season, needle samples will be taken from all individuals (>10,000 samples) and stored for DNA extraction. In their second year of growth, the seedlings will experience either the stress associated with simulated drought conditions (greenhouse trail), or winter hardening in response to the natural seasonal patterns of decreased daylight and temperature (outdoor raised beds). All individuals will be phenotyped for drought and frost tolerance. DNA will be sequenced for all 40 seedlings in each of the 40 wild and 11 orchard seedlots, and from 40 randomly selected seedlings in each of the 20 case-control populations. DNA will also be sequenced for the selected seedlings in the case-control populations that are identified as one of the 10 least or the 10 most tolerant individuals.

In terms of progress, we have finalized the seedlot selections for this experiment. Seedlots were selected to provide adequate climatic and geographic variation across the natural range of coastal and interior Douglas-fir – from Mexico through the western United States into British Columbia and Alberta. The seeds are presently in cold stratification and we are busy making preparations the massive undertaking of filling and sowing 10,000 cones!
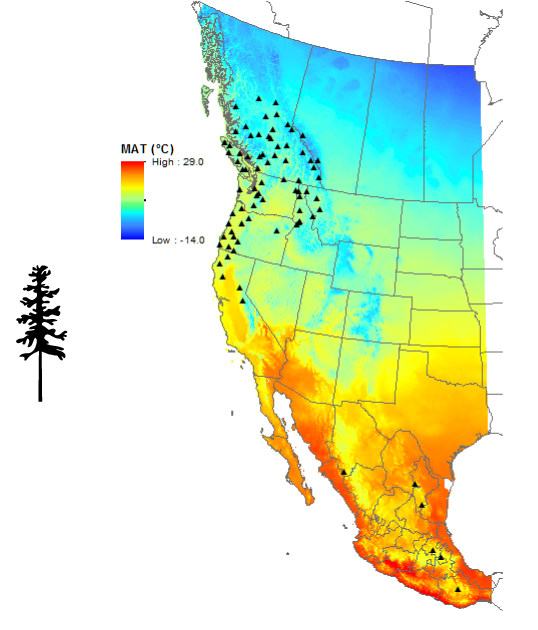
Seed source locations for Douglas-fir.
Activity 1.2 Candidate genes in western larch and jack pine
The comparative genomics for Douglas-fir (Activity 1.1 above) and lodgepole pine (from the AdapTree project) will be used to increase the power for identifying climate-adaptation candidate genes in western larch and jack pine. A genotype x environment interactions association approach will be used to reveal patterns of local adaption to climate for western larch and jack pine. For each species, 30 populations spanning the full geographic and climate range of these species will be sampled from existing provinence trails in British Columbia (western larch) and in Ontario (jack pine). DNA will be extracted from 30 individuals per population. Sequence capture libraries will be made for each population and estimated SNP frequencies will used to detect local adaptation to climate.
To begin this process we have extracted RNA from one actively growing western larch and jack pine seedling. The RNA (presently being sequenced) will be used assemble the de novo transcriptomes.

Western larch shipment arriving at UBC (December 2017) and actively growing in the climate chamber (January 2017).

Jack pine breaking bud (February 2017) and actively growing in the climate chamber (March 2017).
RNA extraction
RNA extractions on the actively growing jack pine and larch were carried out in the lab at UBC in March 2017. RNA sequencing and transciptome assembly will be conducted by the Genome Quebec Innovation Centre at McGill.
To prepare the material for extraction the needle tissue was ground in liquid nitrogen to a fine powder using a mortar and pestle. Total RNA was extracted using Spectrum™ Plant Total RNA kit (Sigma, USA) and cleaned up by the On-Column DNase I Digestion Set (Sigma, USA). The purity of the total RNA preparations were assessed with a NanoDrop spectrophotometer (Thermo Scientific, USA). The concentration was determined by Qubit® 2.0 Fluorometer (Invitrogen, USA). The integrity and size distribution of purified RNAs were evaluated with a 2100 Bioanalyzer (Agilent Technologies, USA) equipped with an RNA Nano chip. For more information on the RNA extraction procedures contact Dragana Obreht Vidakovic.

RNA extraction for jack pine and western larch in the lab at UBC (March 2017).
Activity 2. Disease resistance
Activity 2.1 Dothistroma and lodgepole pine
Activity 2.1.1 Host gene expression in lodgepole pine
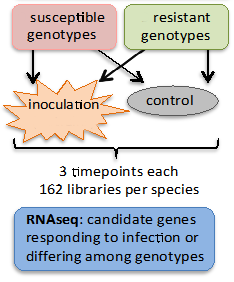 To study gene expression in response to infection we will compare the responses in the progeny of controlled crosses of parents that are highly resistant (resistant x resistant) to those that are highly susceptible (susceptible x susceptible) to Dothistroma. Temporal changes in expression will be monitored at three separate time points during the infection process. RNAseq will be used to characterize differential gene expression, identify candidate genes for resistance and to search for SNPs to be used in the gene wide association study (Activity 2.1.2).
To study gene expression in response to infection we will compare the responses in the progeny of controlled crosses of parents that are highly resistant (resistant x resistant) to those that are highly susceptible (susceptible x susceptible) to Dothistroma. Temporal changes in expression will be monitored at three separate time points during the infection process. RNAseq will be used to characterize differential gene expression, identify candidate genes for resistance and to search for SNPs to be used in the gene wide association study (Activity 2.1.2).
The one year-old, controlled crosses are currently experiencing their first winter in the climate chambers at UBC. When the 54 individuals have formed well-developed second year needles (May-June 2017) we will begin the inoculation procedure.
For more information on the seedlings contact Pia Smets and Christine Chourmouzis.

Germination (December 2016) and budset (February 2017) in lodgepole pine controlled crosses.
Inoculation pilot study
The pilot study designed to develop an effective Dothistroma septosporum (an ascomycete fungus) inoculation procedure for lodgepole pine has been successfully completed! The conidia (the asexual Dothistroma spores), which were cultivated in the lab on a sporulating medium and sprayed on the needles at a concentration of 3 million conidia per ml, achieved a germination success of 95%. The first signs of disease appeared 5 weeks after inoculation and the symptoms after 8 weeks.
For more information Dothistroma innoculation contact Nicholas Feau.

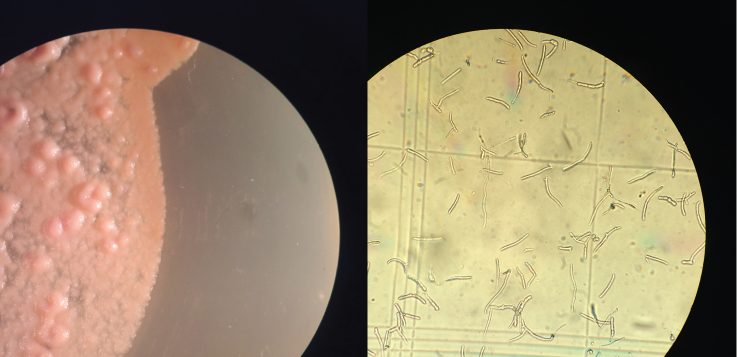
Dothistroma fungus in vitro – fungus on sporulating medium (left) and the conidia (right).

Dothistroma innoculation (left and inset) and incubation under moist conditions (right).

Signs of Dothistroma infection appeared at 5 weeks (left) and symptoms at 8 weeks (right) after inoculation.
Activity 2.1.2 Gene wide association study
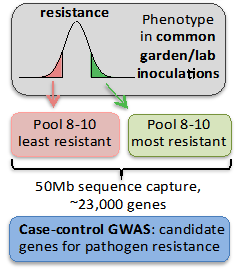 The gene wide association study designed to identify candidate genes and population variation for disease resistance or tolerance to Dothistroma is now in the beginning stage. 100 seeds from each of 40 wild-collected lodgepole pine populations from BC and Alberta have been sown and are presently in the germination phase. The populations selected for this study include pure lodgepole, pure jack pine and several hybrids (lodgepole x jack pine). After innoculation with Dothestroma (fall 2017) phenotypic resistance will be quantitatively accessed in all individuals by using visual measures of infection severity. The 10 least and the 10 most resistant seedlings in each population will be sequenced. The data will be analyzed using the SNP’s generated in Activity 2.1.1 (Host Gene Expression) and used to generate a list of genes with a strong signature of association to resistance phenotypes.
The gene wide association study designed to identify candidate genes and population variation for disease resistance or tolerance to Dothistroma is now in the beginning stage. 100 seeds from each of 40 wild-collected lodgepole pine populations from BC and Alberta have been sown and are presently in the germination phase. The populations selected for this study include pure lodgepole, pure jack pine and several hybrids (lodgepole x jack pine). After innoculation with Dothestroma (fall 2017) phenotypic resistance will be quantitatively accessed in all individuals by using visual measures of infection severity. The 10 least and the 10 most resistant seedlings in each population will be sequenced. The data will be analyzed using the SNP’s generated in Activity 2.1.1 (Host Gene Expression) and used to generate a list of genes with a strong signature of association to resistance phenotypes.
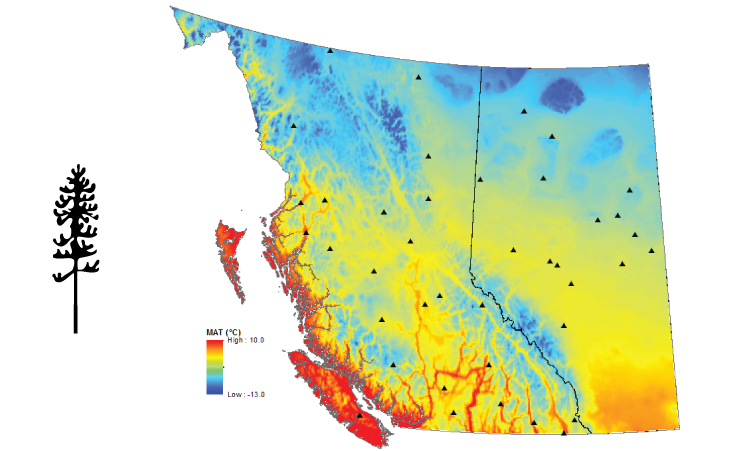
Location of the pine in British Columbia and Alberta.

Seed preparation and sowing (February 2017).

Cone trays in the climate chamber at UBC (March 2017).

Germinating seeds (March 13 2017).
| cell 1 | cell 1 |
Activity 3.
Updates coming soon. We need content here. Who should I contact for this? Any ideas? It might be easier to get something once people see it only needs to be brief and very general in terms of content for these “Highlight” pages.
Activity 4.
It will be a while yet.
Activity 5.
Updates coming soon. I need to speak with Shannon and have her contribute something
