Research Progress HIGHLIGHTS
Activity 1 – Climate adaptation
Activity 1.1 Candidate genes in Douglas-fir
To investigate patterns in the distribution of adaptive phenotypes and to identify candidate genes relating to drought and frost tolerance in Douglas-fir, we have established 1) a large outdoor common garden trial to test frost hardiness, and 2) two large indoor container experiments to examine response to drought stress. We have grown seedlings from 73 wild collected (natural) populations and 14 orchard seedlots (selectively bred trees). Of the 73 wild seedlots, 20 were used for a case-control approach in which genotype-phenotype association analyses (GWAS) will be performed. Genotype-environment association (GEA) tests will be performed for all the 87 sampled populations. Seven of the 14 orchard seedlots are part of a separate study on assisted migration – these seven seedlots are presently growing on 45 different sites (i.e. climates) across western Canada and the US where they are being phenotyped for survival and growth.
Seedlot selection was finalized in December 2016. The seedlots were chosen to provide adequate climatic and geographic variation across the natural range of coastal and interior Douglas-fir. They span a range of more the 16° latitude, 14° longitude and from 61 m to 1737 m in elevation.
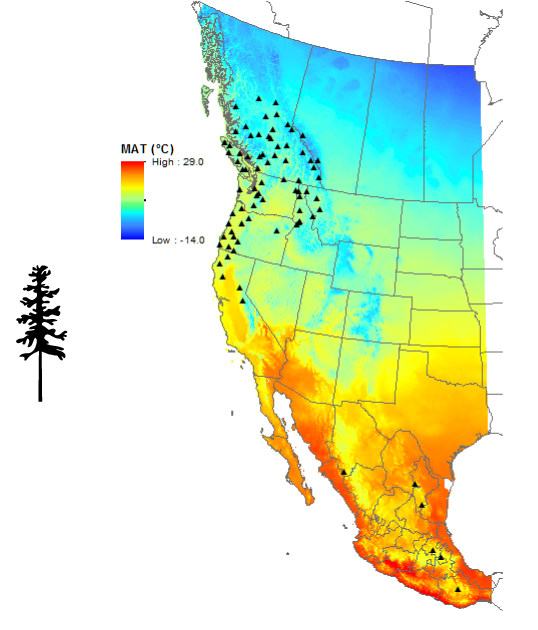
Map of seed source locations for Douglas-fir.
The seeds were stratified in April 2017, double sown into small cones at the greenhouse at UBC in May 2017 and thinned in June 2017. At the end of their first growing season, needle samples were taken from all individuals (7,200 samples) and stored for subsequent DNA extraction. For each of the 87 provenances 40 seedlings were randomly chosen and their DNAs were extracted, quantified and subsequently pooled for sequence capture (for the GEA).

Year 1. Douglas-fir germinates (May 2017) and the discards after thinning the double sowed seedlings (June 2017).
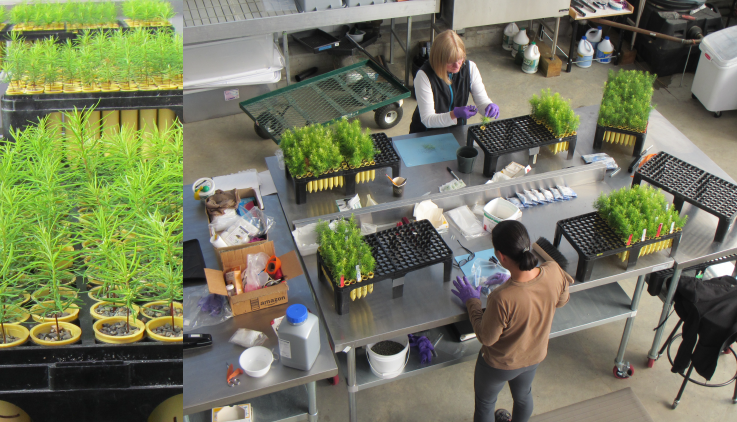

Year 1. Douglas-fir seedlings (July 2017) and tissue sampling for DNA extraction (October 2017).
In their second year of growth, the seedlings were divided in two experimental groups: frost and drought. In March 2018, seedlings in the frost group were planted into large, deep, outdoor raised beds, where they were exposed to the natural seasonal patterns of daylight length and temperature. They were measured throughout the two subsequent growing seasons (e.g. growth, bud burst, but set timing), tested for cold hardiness in the fall of 2018, and harvested for root collar diameter and shoot biomass measurements at the end of the growing season of 2019. Approximately 160,000 phenotypic observations were generated from this experiment.
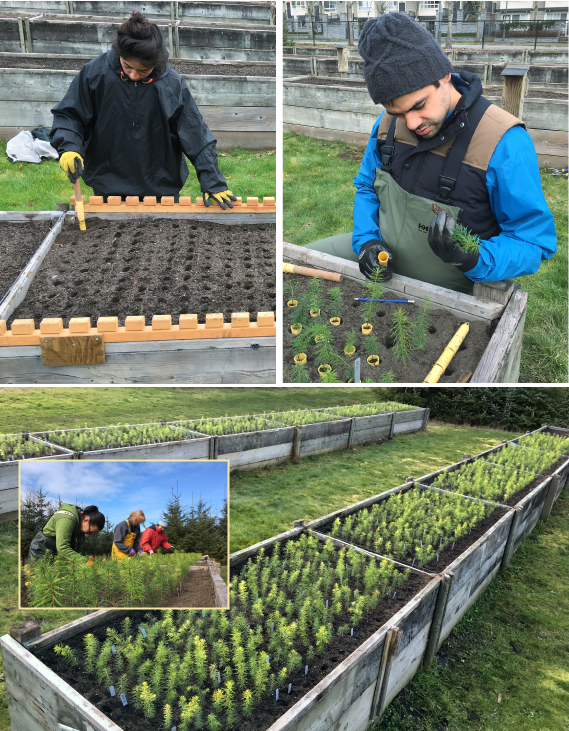
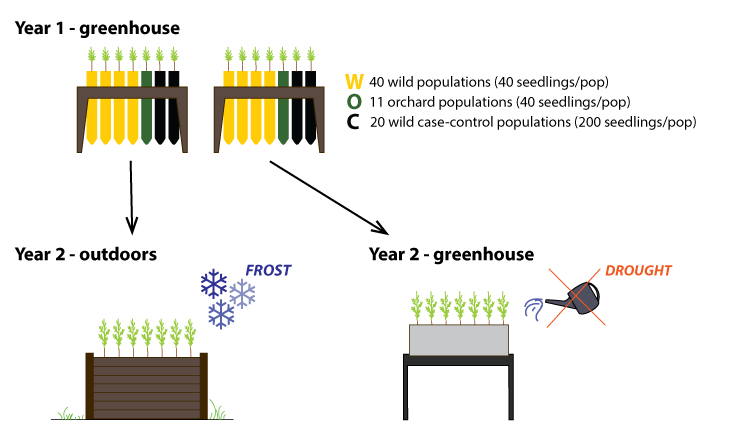

Planting Douglas-fir seedlings in the outdoor raised beds (March 2018).
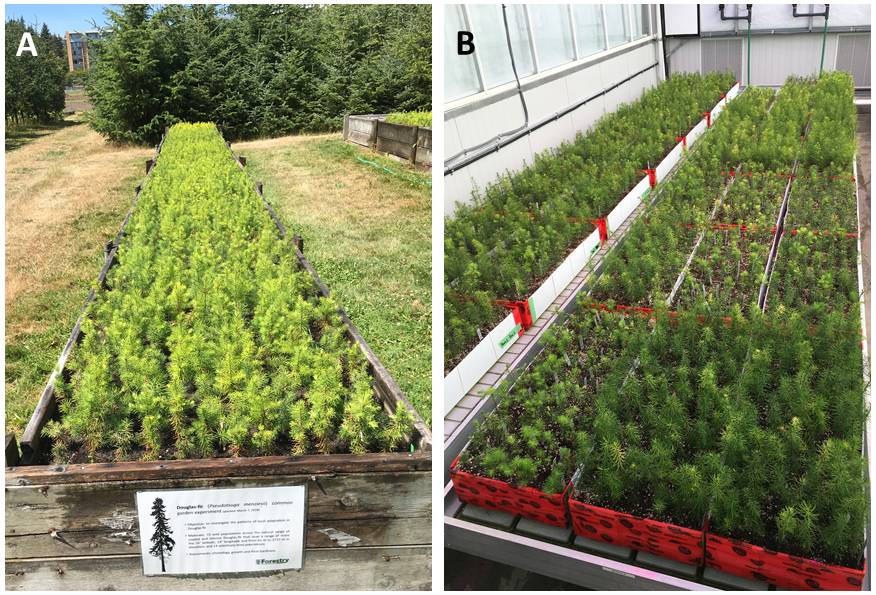

A Cold-hardiness experiment planted in raised beds, UBC Totem Field experimental station.
B Drought experiments 1 and 2 (right to left), planted in multi-seedling containers, UBC Botany greenhouse.
In April 2018, the remaining seedlings were planted into multi-seedling containers for the drought tolerance experiments. Two separate experiments were established for drought-hardiness phenotyping. The first experiment (“drought 1”) included only the 20 case-control seedlots with 74 to 80 seedlings from each. The second experiment (“drought 2”) was established with 4 to 18 seedlings per seedlot from all populations (87 seedlots). Seedlings spent 8 weeks in these containers under optimal conditions for acclimation and to start their active growth.
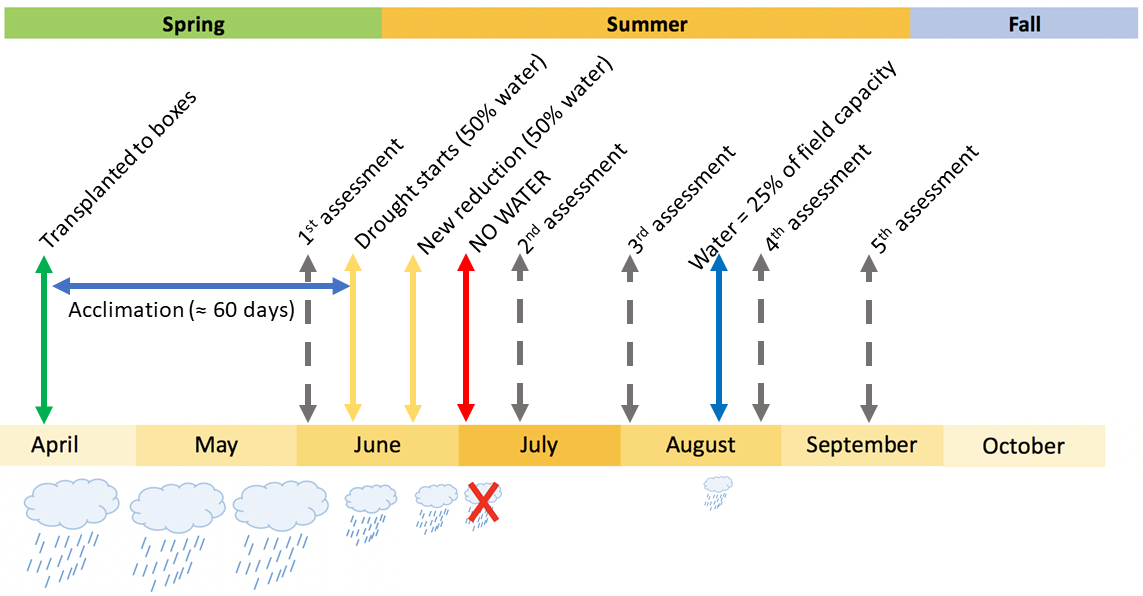
Drought 2 schema with the watering regime and the assessments.
The drought treatment, which started in June 2018 and was applied in both experiments after the acclimation period, involved reducing watering gradually until the soil reached moisture levels below the expected wilting point. Over the following weeks the target moisture in the soil was a predawn water potential below -1.5 MPa and a volumetric water content below 10%.
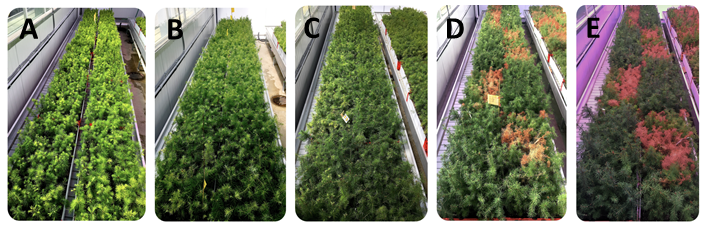

Drought 2 at different stages after treatment: A – May, B – June, C – July, D – August, E – September.
Phenotyping of seedlings for the drought experiments consisted of recording survival, visual drought damage, growth and photosynthetic efficiency (chlorophyll fluorescence) systematically through time in all tested seedlings. Approximately 58,500 phenotypic observations were generated from both drought experiments combined.
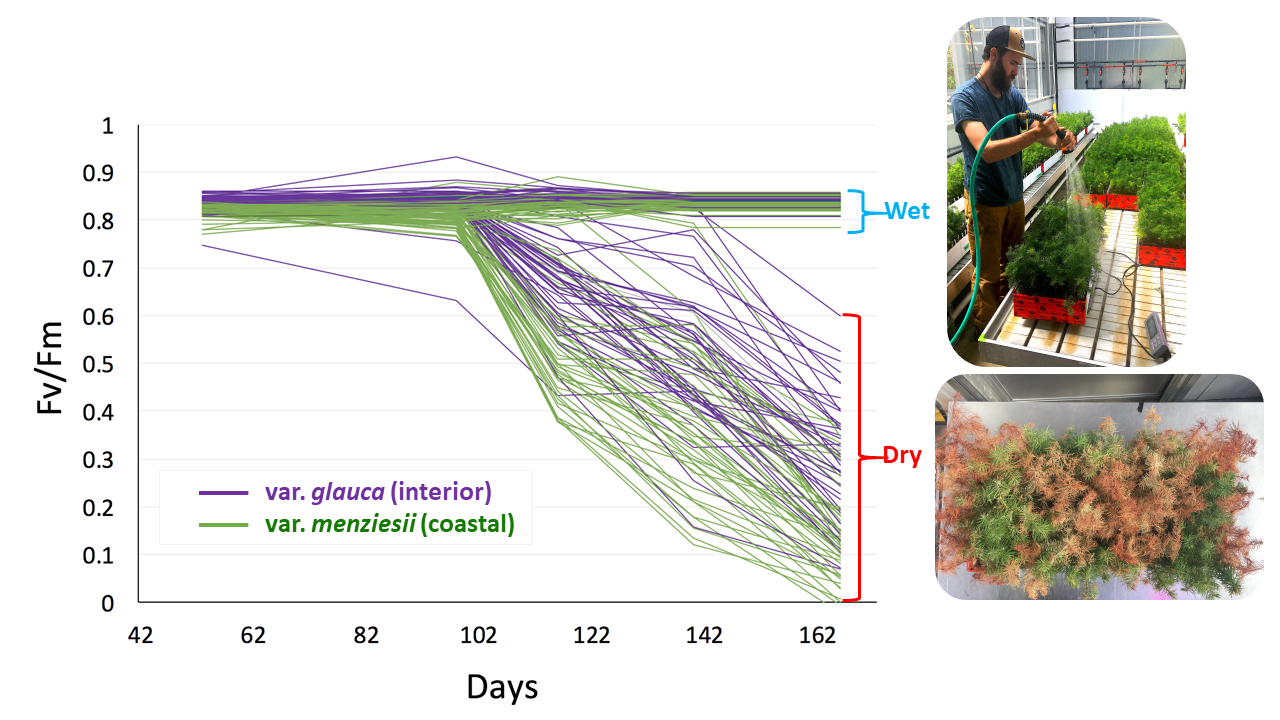
Drought 2 progress of average Fv/Fm through time for each population (after correction for spatial autocorrelation). Green lines depict var. menziesii populations and purple lines depict var. glauca populations.
For the 20 case-control populations in the drought and frost experiments, we selected the 10 least and the 10 most tolerant individuals to be sequenced. In the frost-hardiness experiment, the 10 most hardy individuals per provenance were selected based on the lowest indexes of injury caused by the coldest temperature tested in each variety (-8˚C for var. menziesii and -13˚C for var. glauca); and the 10 least hardy individuals based on the highest indexes of injury caused by the mildest temperatures (-6˚C for var. menziesii and -9˚C for var. glauca).

Frost – example of pooling after selection within one of the var. glauca case-control provenances (provenance 3). In each tested temperature (-9˚C and -13˚C) each point represents one individual of the provenance. Blue points represent the most frost-hardy individuals and red points the least hardy ones. The red and blue circles indicate the pooled case and control groups. Error bars depict the 95% confidence intervals.
The 10 most and 10 least drought hardy individuals from each of the 20 case-control provenances (10 var. menziesii and 10 var. glauca) were selected based on the decay of chlorophyll fluorescence (decrease in photosynthetic performance) through time on after extreme drought.
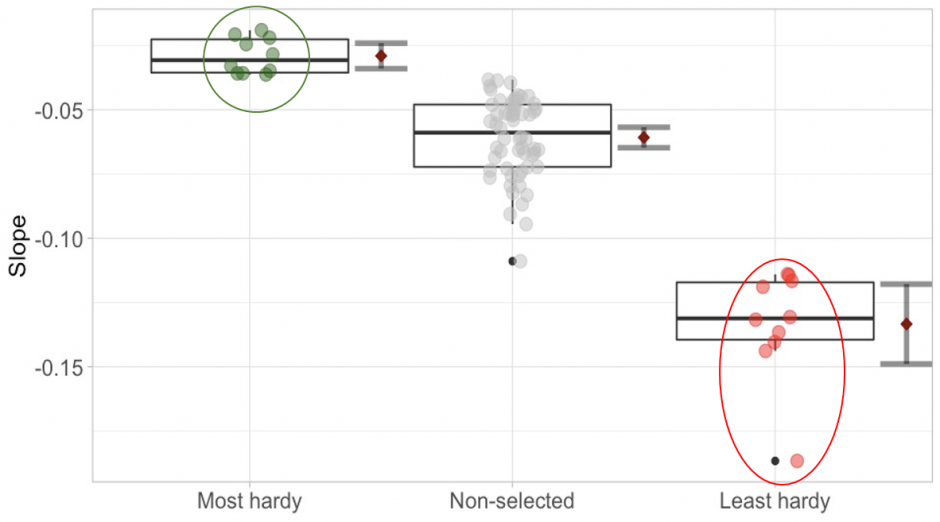
Drought 1 – example of pooling after selection within one of the var. glauca case-control provenances (provenance 3). Each point represents one individual of the provenance. Green points represent the most drought-hardy individuals and red points the least hardy ones. The green and red circles indicate the pooled groups of selected individuals. Error bars depict the 95% confidence intervals.
After selection, individuals’ DNAs in each group of 10 seedlings (10 most or 10 least hardy) from each experiment (drought and frost) and from each of the 20 case-control populations, were extracted, quantified, and subsequently pooled for sequence capture (for the GWAS).
We are now establishing the protocols for putting together the phenotype data with the sequencing data, and for performing the GWAS and the GEA analyses. These analyses will provide us with information to tackle the genetic variants underpinning complex adaptive traits that are related to the cold and drought hardiness as well as help us to find the most important climatic variables determining the Douglas-fir local adaptation.
For more -information, contact Rafael CandidoRibeiro
Activity 1.2 Candidate genes in western larch and jack pine
The comparative genomics for Douglas-fir (Activity 1.1 above) and lodgepole pine (from the AdapTree project) will be used to increase the power for identifying climate-adaptation candidate genes in western larch and jack pine. The genomics of climate adaptation in Douglas-fir will be characterized in comparison with western larch (Pseudotsuga and Larix are sister genera) and jack pine will be compared to lodgepole pine. Additionally, there will be a four-way contrast among all species. The extent of parallel adaptation among conifer species can provide a window into the genomics of complex traits and increase the power to separate false positives from true climate-driven genetic patterns.
A genotype x environment interactions association approach will be used to reveal patterns of local adaption to climate. Sequence capture method will be applied to enrich the exonic regions for each species. Sequence capture method combines the hybridization capture and high-throughput sequencing, allowing the rapid and simultaneous description of numerous and potentially large targeted DNA regions. Capture probes are used to target and hybridize the DNA library. We can estimate SNP frequencies from the sequenced target regions.
Capture probes
For designing capture probes, we began by assembling de novo transcriptomes for jack pine and western larch. The transcriptome is the set of all RNA molecules in one cell or a population of cells. De novo transcriptome assembly is the de novo sequence assembly method of creating a transcriptome without the aid of a reference genome. We extracted RNA from foliage of one jack pine and one western larch seedling and sequenced the RNA. The assembled transcripts were then used to design a ~50 Mb sequence capture probe panel for each species.
RNA extraction
RNA extractions on the actively growing jack pine and larch were carried out in the lab at UBC in March 2017. RNA sequencing and transciptome assembly will be conducted by the Genome Quebec Innovation Centre at McGill.
To prepare the material for extraction the needle tissue was ground in liquid nitrogen to a fine powder using a mortar and pestle. Total RNA was extracted using Spectrum™ Plant Total RNA kit (Sigma, USA) and cleaned up by the On-Column DNase I Digestion Set (Sigma, USA). The purity of the total RNA preparations were assessed with a NanoDrop spectrophotometer (Thermo Scientific, USA). The concentration was determined by Qubit® 2.0 Fluorometer (Invitrogen, USA). The integrity and size distribution of purified RNAs were evaluated with a 2100 Bioanalyzer (Agilent Technologies, USA) equipped with an RNA Nano chip. For more information on the RNA extraction procedures contact Dragana Obreht Vidakovic.

RNA extraction for jack pine and western larch in the lab at UBC (March 2017).

Western larch shipment arriving at UBC (December 2017) and actively growing in the climate chamber (January 2017).

Jack pine breaking bud (February 2017) and actively growing in the climate chamber (March 2017).
Sequencing libraries
For constructing sequencing libraries, we sampled foliage/bark from existing range-wide western larch and jack pine provenance trials and seed collections. For each species, we selected 40 populations, with 20 individuals, that span the full geographic and climatic range of each species. The DNA is presently being extracted from each individual with representation equalized in pools of all individuals for each population. In spring 2018, we will construct capture libraries for each population by hybridizing the probes to the pooled DNA libraries. After the capture libraries are sequenced, we will estimate SNP frequencies for each population from reads in sequenced population pools and apply the analytical approaches developed in the previous project for genotype-environment association.
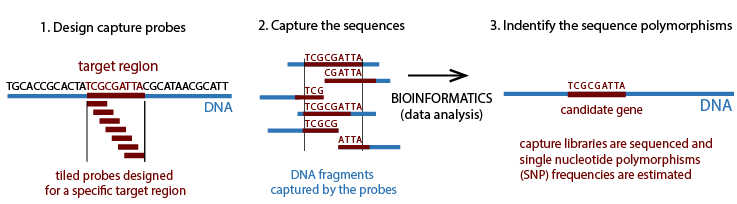
We will use sequence capture probes (~50 Mb sequence panels) to capture genomic regions of interest in jack pine and western larch.
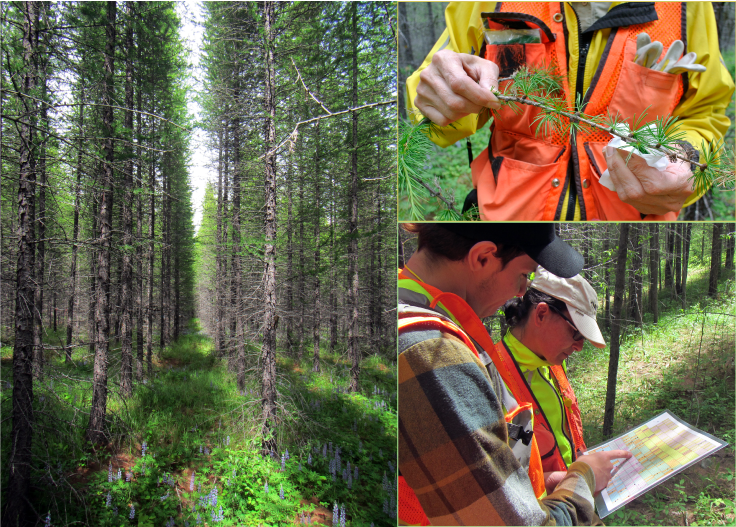

Collecting western larch needles in the Lamb Creek BC provenance trial (June 2017).
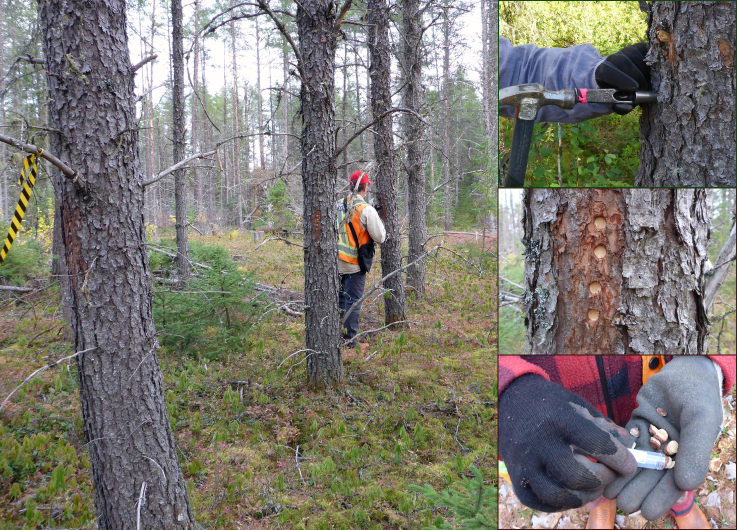 Bark sampling in jack pine provenance trials (Sept 2016 and July 2017).
Bark sampling in jack pine provenance trials (Sept 2016 and July 2017).
For more information, contact Mengmeng Lu and Sam Yeaman.
Activity 1.4 Comparative transciptomics of Douglas-fir
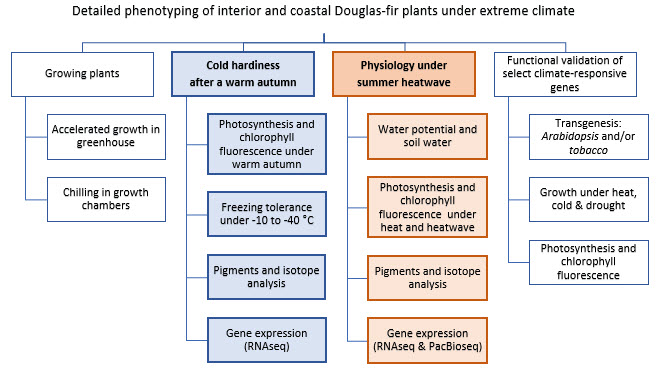
How do trees adapt to different climates? In this activity, we are comparing the physiological responses of four Douglas-fir provenances from different home environments (two coastal and two interior) to simulated future dryer and warmer climates. We will zoom in on gene expression (RNA, the transcriptome) to identify climate-responsive genes. The physiological mechanisms being studied include photo-protection, energy dissipation, water-use efficiency, and photosynthetic gas exchange. This study will result in a more complete causal description of abiotic stress response in Douglas-fir.
We have grown two batches of Douglas-fir seedlings. The first batch has gone through an accelerated growing cycle and are now being used to study the effects of warm autumn to the development of cold hardiness (started in December 2017). The second batch of seedlings are now in their second growing season and will be used to study the response to heatwave (set for March 2018). Using transcriptome data and detailed phenotyping of Douglas-fir responses to extreme climates, we will select a few climate-responsive genes. These candidate genes will be functionally validated using model plants.
Warm Autumn – Effects of a warmer climate on late season performance and the development of cold hardiness.
Douglas-fir seedlings exposed to a short photoperiod (needed to initiate budset) and summer temperature will be compared to seedlings exposed to normal autumn temperature. The difference in photosynthesis and chlorophyll fluorescence will be measured using a portable gas exchange fluorescence system and several pigments will be quantified to understand photoinhibition in these plants. Both sets of plants will be exposed to below freezing conditions to determine their level of freezing tolerance. The phenotypic data will be complemented by RNA-seq quantification of gene expression in needles, buds and roots.
Heatwave – Differences in the response of Douglas-fir provenances to heatwave.
In this experiment, we will subject Douglas-fir plants to seven days of no water supply, followed by up to two weeks of extremely hot days. We will compare the responses of heatwave treated Douglas-fir to plants grown under extreme hot and normal summer conditions. Water potential and soil water will be monitored regularly and photosynthesis and photoinhibition will be measured weekly during the treatment. Samples will be collected for quantification of pigments, carbon stable isotope and gene expression profiling.
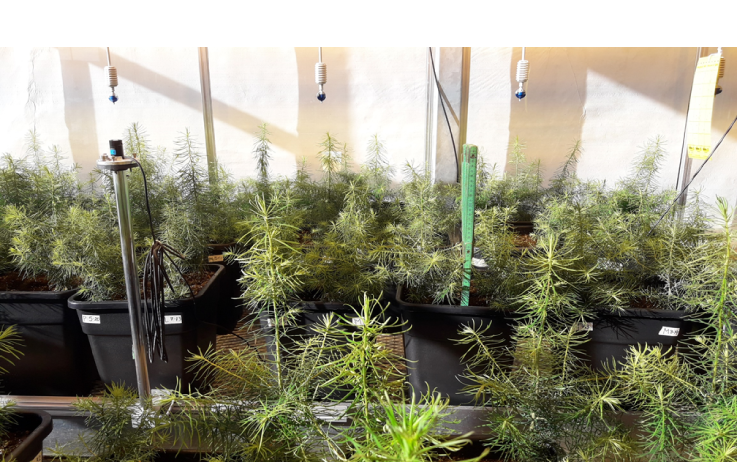

Douglas fir seedlings growing in the research greenhouse at University of Toronto Mississauga. These seedlings are in their second growing season and will be used to study response to heatwave in March 2018.
For more information, contact Vera Velasco and Ingo Ensminger
